The emerging world of rapid tests
Rapid diagnostics for faster medical decision making
Healthcare relies heavily on laboratory results to help make diagnostic and treatment decisions.
Therefore, laboratory tests must be of high quality and have adequate turnaround times.
Timely and reliable microbiological diagnostics are needed to select the most appropriate antibiotics or antibiotic combinations for patients.
The answers to the following questions provide information to clinicians to make a therapeutic decision:
- Is it a bacterium or not?
- What type of bacterium?
- What is the bacterium sensitive/resistant to?
Is it a bacterium or not?
By measuring the amount of C-reactive protein (CRP) in the blood, we can obtain information about the extent of inflammatory processes in the body. A low level of CRP indicates that there is no bacterial infection, and therefore antibiotic treatment is not required.
(Source of image: https://commons.wikimedia.org/wiki/File:1lj7.jpg)
What type of bacterium?
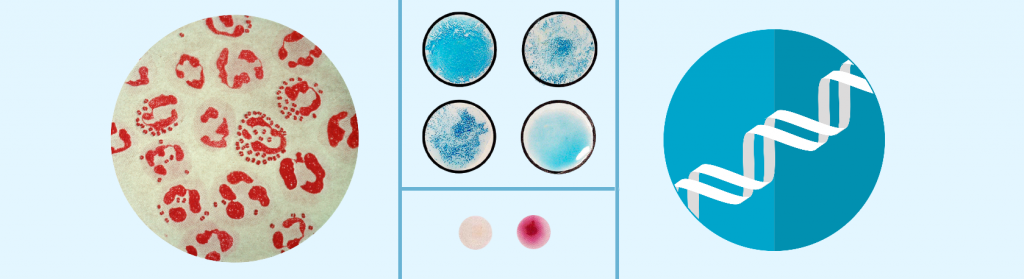
Knowing the name of the bacterium or only which larger group it belongs to can already give us information on how to treat it.
Several rapid tests are available to identify bacteria:
⇾ Direct microscopic examination: Gram staining
The first step in diagnostic processing is generally a direct microscopic examination. With the Gram stain, most bacteria and also inflammatory cells can be quickly visualised. Its diagnostic value is high in soft tissue abscess, lower respiratory tract infection, urinary tract infection, cervicitis, and urethritis. It also provides help with wound exudate as a preliminary result.
⇾ Antigen-detection assays / agglutination test
Streptococcal group identification based on Lancefield antigens is a common test in microbiological laboratories, especially the Group A beta-hemolytic Streptococcus, the Streptococcus pyogenes. Accurate and rapid diagnosis of S. pyogenes is important as there is a possibility that throat and skin infections could lead to severe life-threatening invasive conditions and post-infection immune-mediated complications if left untreated.
⇾ Rapid biochemical tests
Biochemical tests help identify bacteria based on differences in their biochemical activity.
For example, the coagulase test is used to distinguish between Staphylococci or the PYR-test that detects the pyrrolidonyl arylamidase enzyme is used for the rapid differentiation of enterococci from group D streptococci, which have different antibiotic sensitivities.
⇾ Molecular detection and identification
Some of these methods can provide results in a couple of hours and are suitable not only for the identification of bacteria but also for the determination of resistance:
- polymerase chain reaction (PCR)
- loop-mediated isothermal amplification (LAMP)
- matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) analysis
- peptide nucleic acid fluorescent in situ hybridization (PNA-FISH)
- a blood culture nanotechnology microarray system for Gram-negative bacteria (BC-GN), and for Gram-positive bacteria (BC-GP)
- other new approaches: e.g. single-cell methods, microfluidic approaches
What is the bacterium sensitive/resistant to?
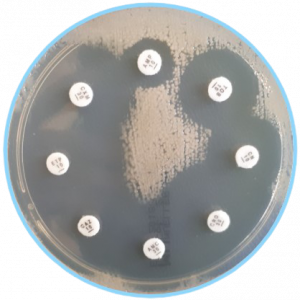
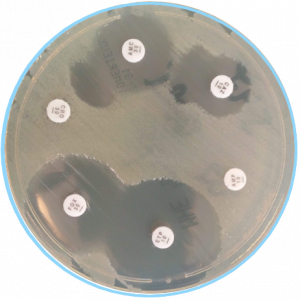
⇾ Detection of phenotypic resistance
Bacteria are tested for growth in the presence of antimicrobial agents.
To these methods, isolated bacteria are required, and isolation is time-consuming.
These methods provide information for clinical management and surveillance but not direct information about the mechanism(s) of resistance to the agent.
- disk diffusion
- combination disk test
- Etest – determine the minimum inhibitory concentration (MIC) – the gold standard.
⇾ Molecular methods for AMR
Molecular AMR diagnostics detect resistance-coding genes or resistance-associated mutations in DNA extracted from purified bacterial isolates or directly from clinical samples.
Characterisation of resistance genes provides essential information for a better understanding of AMR molecular epidemiology.
- amplification tests: PCR and LAMP
- hybridisation tests: FISH and LPAs (line probe assay)
- immunoassays: LFIA (lateral flow immunoassay)
Importance of rapid detection of resistance
The current workflow for detecting the resistance in the microbiological laboratories mostly requires the isolation of the bacteria and the application of different methods.
These techniques take between 16-30 hours, meaning that the appropriate antibiotic treatment is delayed.
Rapid detection is essential to prevent further spread and fight efficiently against MDR bacteria. In some cases rapid detection is lifesaving: the bacterial blood infections that can lead to sepsis are particularly time-sensitive. For this indication, every hour that infection remains untreated with effective antibiotics decreases the patient’s survival rate by 7.6%.
The situation is similar for ventilator-associated pneumonia (VAP) and complicated urinary tract infections.
Significant time can be saved by direct AMR detection from a patient sample.
Quick screening prior to and during admission to the hospital in non-emergency settings – especially within oncology, haematology, neonatal, and intensive care units – can help doctors better detect the type of drug-resistant bacteria present within patients (if any) and provide guidance for treatment.
AMR DetecTool

Affordable, sensitive, rapid and easy to use diagnostic system to detect multidrug-resistant bacteria direct from clinical samples: blood culture, urine, rectal swab, bronchoalveolar lavage / tracheal aspirate.
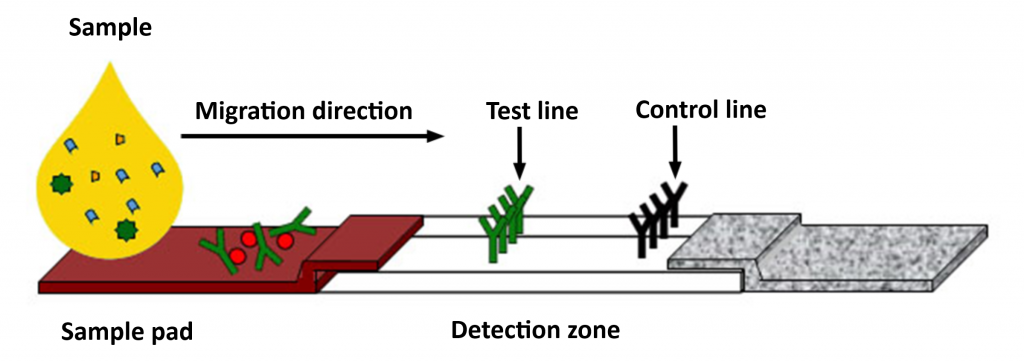
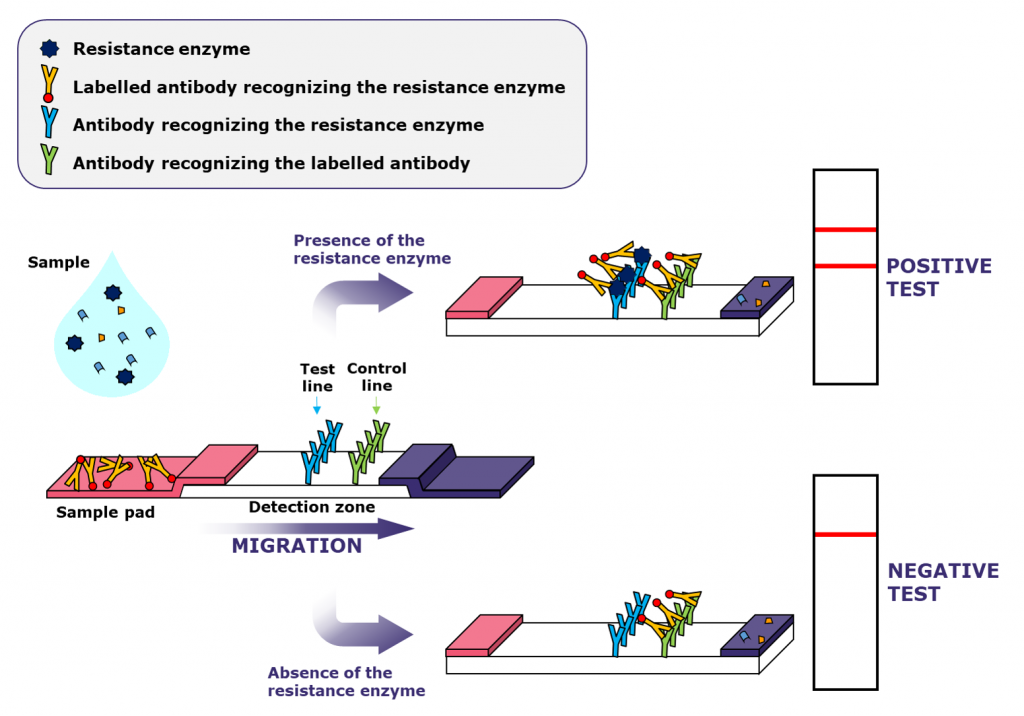
Principle of the detection: LFIA
Key features of the AMR DetecTool:
- Concentrates bacteria from a liquid sample
- Eliminates the interferences due to the biological media
- Improved sensitivity
In addition to detecting resistance, the device also provides sample processing: filtration – extraction – incubation – deposition on the strip.
The sample is applied at the end of the strip (sample pad), then migrates into the detection zone. If the target is present in the sample, the antibodies will bind to it, and coloured bands will appear.
Sources:
https://www.mlo-online.com/home/article/13002427/rapid-diagnostic-testing-in-microbiology
https://www.sciencedirect.com/science/article/pii/S0361112478801282?via%3Dihub
https://apps.who.int/iris/bitstream/handle/10665/310993/WHO-WSI-AMR-2019.1-eng.pdf?ua=1
https://academic.oup.com/jac/article/73/4/909/4819243
https://journals.sagepub.com/doi/full/10.1177/2211068216680207

Copyright 2020 – AMRDETECTOOL
